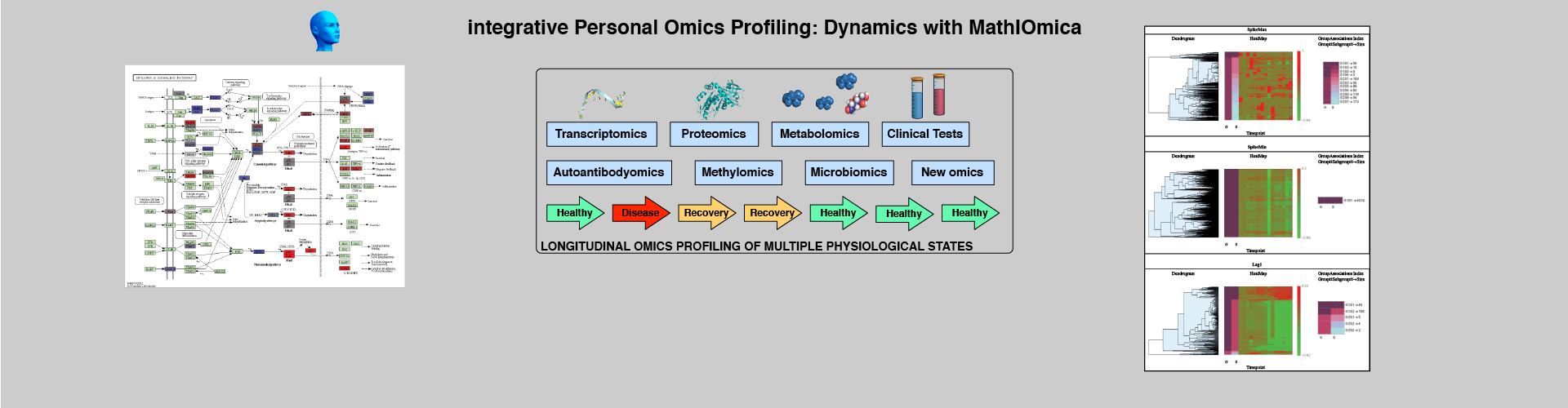
Time Series Evaluation and Classification
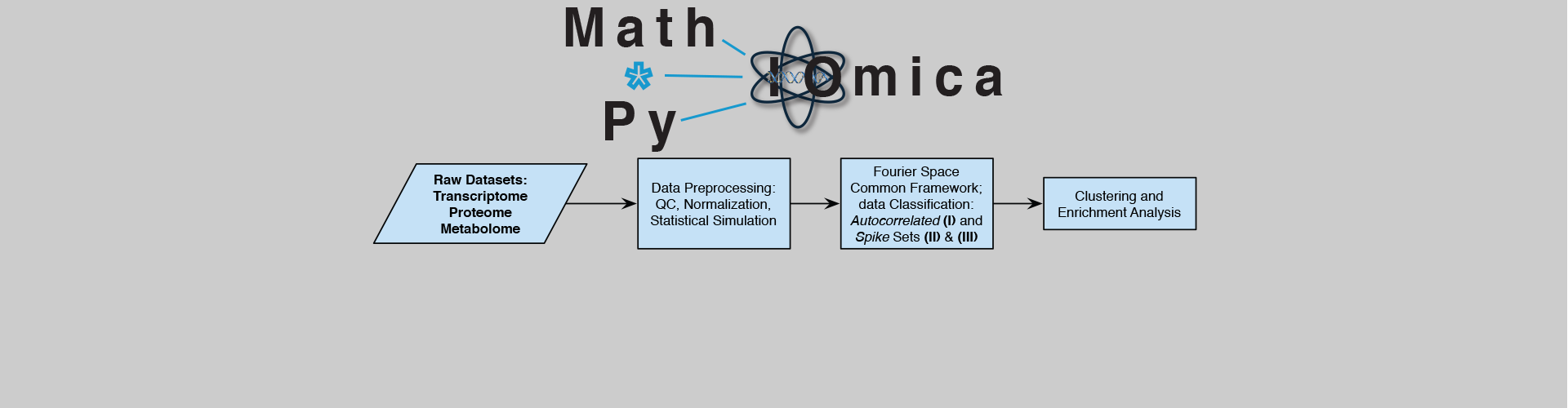
Data Integration with MathIOmica and PyIOmica.
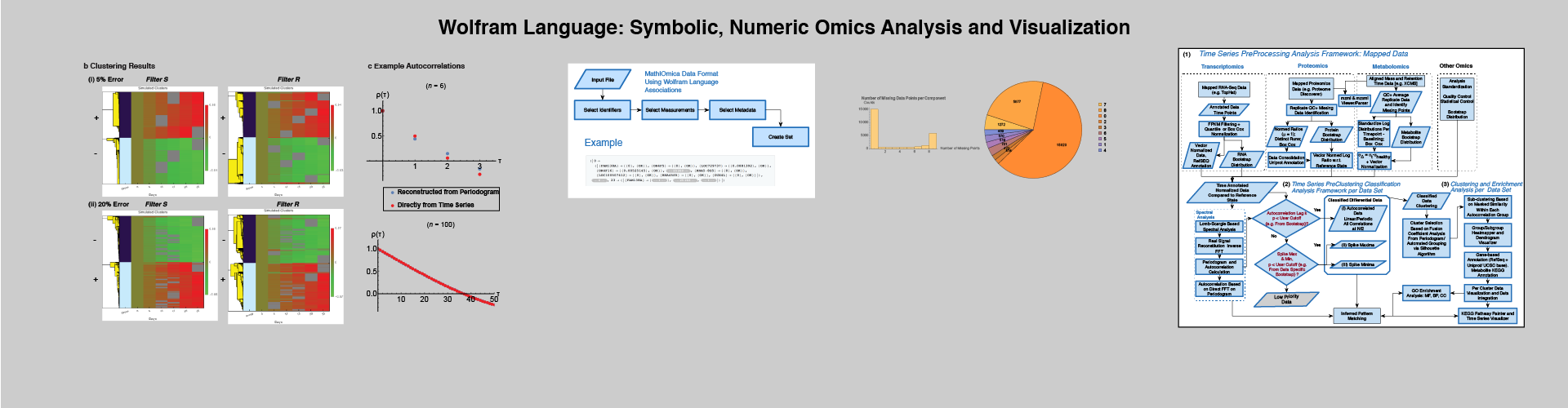
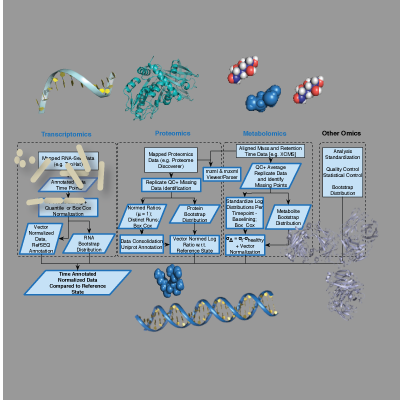
Extensible Framework
MathIOmica is based on the Wolfram Language and PyIOmica is written in Python. These *IOmica programs provide a framework for graphical, numerical and symbolic work for omics analyses. The code is cross-platform, open source and includes full integrated documentation.
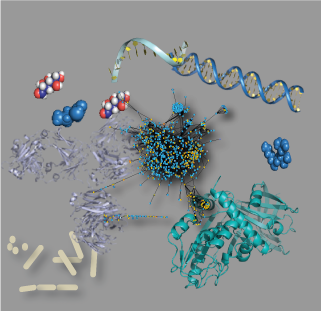
Omics analyses and integration require extensive processing. MathIOmica and PyIOmica provide a user-friendly framework for downstream analysis and visualization.
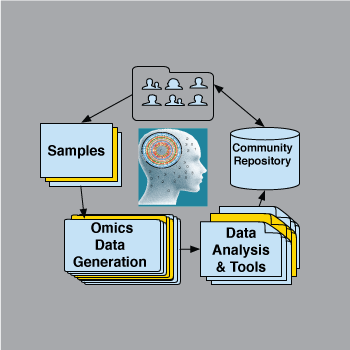
Creating models needs robust omics datasets, particularly for time series analyses and network inference. We are building multiple such sets for the community.
*IOmica: MathIOmica and PyIOmica offer unique platforms for omics analysis and integration.
See also
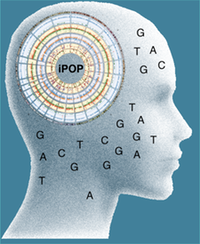
"Personalized medicine is expected to benefit from combining genomic information with regular monitoring of physiological states by multiple high- throughput methods. Here, we present an integrative personal omics profile (iPOP), an analysis that combines genomic, transcriptomic, proteomic, metabolomic, and autoantibody profiles from a single individual over a 14 month period. Our iPOP analysis revealed various medical risks, including type II diabetes. It also uncovered extensive, dynamic changes in diverse molecular components and biological pathways across healthy and diseased conditions. Extremely high-coverage genomic and transcriptomic data, which provide the basis of our iPOP, discovered extensive heteroallelic changes during healthy and diseased states and an unexpected RNA editing mechanism. This study demonstrates that longitudinal iPOP can be used to interpret healthy and disease states by connecting genomic information with additional dynamic omics activity."
From Personal Omics Profiling Reveals Dynamic Molecular and Medical Phenotypes
Cell, Volume 148, Issue 6, 1293-1307, 16 March 2012
Curated mapped subsets of the pilot iPOP data are utilized in MathIOmica for examples and documentation. The original raw data for the pilot iPOP study have been made publically available as follows:
snyderome contains local repository of iPOP data
PEPTIDE ATLAS
http://www.peptideatlas.org/PASS/PASS00062
Tools available on the gmiaslab GitHub repository
MathIOmica utilizes in-built Mathematica Documentation. A printout of various documentation is provided below in pdf form.
You can also download previous releases from MathIOmica's github page.
Mapped RNA-Sequencing Data
Mapped Proteomics Mass Spectrometry Data
Aligned Metabolomics Mass Spectrometry Data
Data from multi-omics analysis (Raw and mapped) will be made available (Mias, Im et al. in preparation).
Mass Spectrometry Spectral Viewer, MathIOmica-MSViewer